FRIENDS OF THE
HEBREW UNIVERSITY
News & Updates
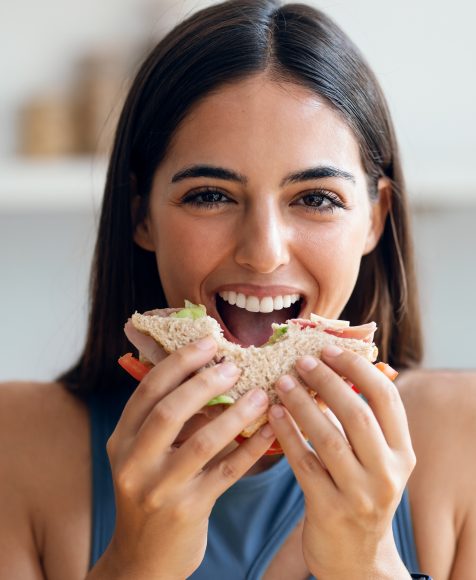
Study Suggests Training Reduces Anxiety for Women with Restrained Eating Habits, Allowing Enjoyment of High-Calorie Foods
April 22, 2024 — Women who are restrained eaters to control their weight can still appreciate high-calorie foods with training to respond more flexibly to food cues, according to a new study by Hebrew University of Jerusalem researchers published in the Journal of Food Disorders. Restrained eaters
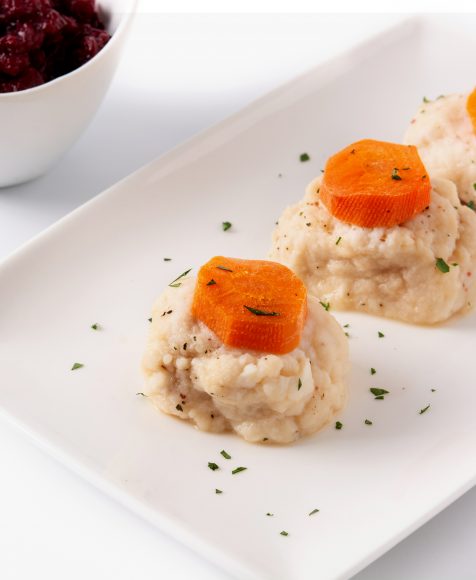
Disease-Resistant Carp: Enhancing Gefilte Fish Quality
April 18, 2024 — According to a new study led by Prof. Lior David from the Faculty of Agriculture at the Hebrew University of Jerusalem focusing on the common carp—a widely produced fish in the aquaculture industry and often the main ingredient in the Passover dish, gefilte fish—selective
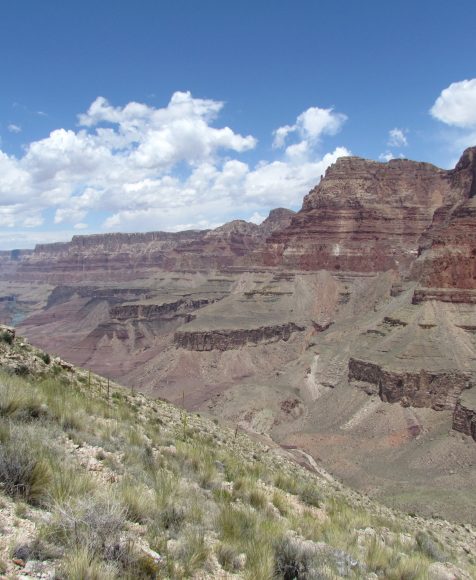
New Approach that Reveals the Evolution of Ancient Ocean Oxygenation Developed by Hebrew University and U.S. Researchers
April 12, 2024 — A new approach to reconstruct the increase of oxygen in ancient marine environments using Uranium-lead (U-Pb) dating of dolomite rocks to detect signals of oxygenation has been developed by a team of researchers led by the Hebrew University of Jerusalem. The findings have
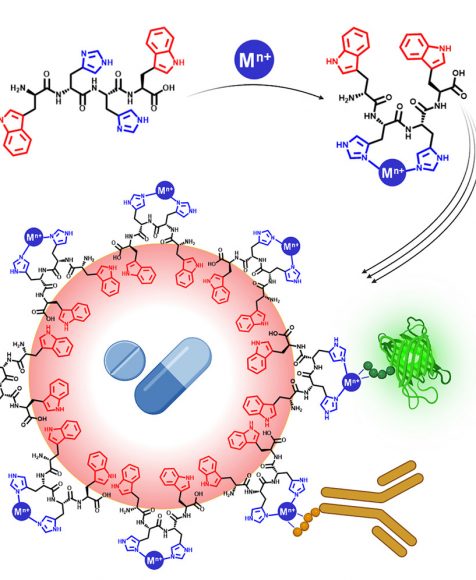
New Drug Delivery System Developed by Hebrew University Researchers Simultaneously Delivers Diverse Therapeutic Agents in a Single Carrier
April 12, 2024 — A new drug delivery system that overcomes previous limitations in delivering diverse therapeutic compounds in a single carrier has been developed by Hebrew University of Jerusalem researchers. According to the new study published in Cell Press, the innovative drug delivery system
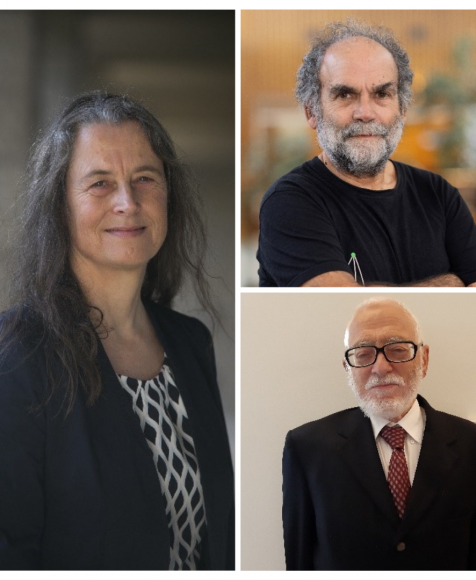
Three Hebrew University Researchers Awarded Prestigious Research Grants by the European Research Council
April 12, 2024 — The Hebrew University of Jerusalem (HU) is thrilled to announce that three of its esteemed faculty members, Professor Maren R. Niehoff, Professor Nathan (Nati) Linial, and Professor David Kazhdan, have been awarded prestigious research grants by the European Research Council
The important thing is to not stop questioning.
Curiosity has its own reason for existing.- Good ol' Al
Upcoming Events
Events are a great way to meet, mingle, and network with luminaries, philanthropists, and visionaries who share a passion for knowledge.
05/09/2024 12:00 PM
54th Annual George A. Katz Torch of Learning Award Luncheon
05/16/2024 6:00 PM
We Are One: An Evening of Inspiration and Solidarity with AFHU
06/01/2024
87th International Board of Governors

Hebrew University at a Glance
484 Awards of Excellence
Including 8 Nobel Prizes in Chemistry, Physics, and Economics.
3,400+ Research Projects
Representing 30% of all academic scientific research in Israel.
10,750+ registered patents
Covering 2,800 inventions, more than 900 licensed technologies, and the launch of 170 spin-off companies.
25,000
Students
Studying across 6 campuses, 7 faculties, together with 1,300 researchers.
JOIN OUR MAILING LIST
Subscribe to hear the latest and greatest from HU.

SUPPORT THE UNIVERSITY
Donations help ensure that the Hebrew University of Jerusalem continues to be Israel’s foremost institution of academic excellence and research.
PLANNED GIVING
Leave a legacy through planned giving while honoring one of humanity’s most noble traits: the quest for knowledge. Planned giving is a powerful way to secure the future for the next generation—and includes the additional perk of tax benefits for you and your loved ones.
Special Initiatives
AFHU LEAD
AFHU LEAD identifies, cultivates, and mentors the next generation of leaders for American Friends of the Hebrew University (AFHU). By engaging seasoned facilitators and brilliant Hebrew University researchers, AFHU LEAD connects future leaders to Israel’s premier research university, the Hebrew University of Jerusalem.

Rimon Initiative
The Rimon Philanthropic Investment Initiative is a new, philanthropy-based venture creation engine from AFHU in partnership with the Hebrew University of Jerusalem; Yissum, the university’s technology transfer company; and ASPER HUJI-Innovate, the university’s center for innovation and entrepreneurship.
Igniting Impact
Supporting Social Initiatives at HU, is a new campaign from AFHU designed to propel social impact initiatives with a global outreach. By selecting the project that speaks to you, you will be able to identify, support, and follow your philanthropic impact.
AFHU LEAD
AFHU LEAD identifies, cultivates, and mentors the next generation of leaders for American Friends of the Hebrew University (AFHU). By engaging seasoned facilitators and brilliant Hebrew University researchers, AFHU LEAD connects future leaders to Israel’s premier research university, the Hebrew University of Jerusalem.
Rimon Initiative
The Rimon Philanthropic Investment Initiative is a new, philanthropy-based venture creation engine from AFHU in partnership with the Hebrew University of Jerusalem; Yissum, the university’s technology transfer company; and ASPER HUJI-Innovate, the university’s center for innovation and entrepreneurship.
Igniting Impact
Supporting Social Initiatives at HU, is a new campaign from AFHU designed to propel social impact initiatives with a global outreach. By selecting the project that speaks to you, you will be able to identify, support, and follow your philanthropic impact.