December 1, 2023 — In a pivotal discovery, Hebrew University of Jerusalem researchers have unraveled the mechanisms governing how bacteria spores retain vital genetic information during dormancy, and activate when revived.
The study, published in Molecular Cell, highlights the discovery of a central chromosomal domain within dormant spores and was spearheaded by Prof. Sigal Ben-Yehuda of the Hebrew University Department of Microbiology and Molecular Genetics and her team.
Understanding this process not only sheds light on bacterial survival in harsh conditions but could offer insights into sustaining long-term transcriptional programs in diverse organisms. Such knowledge might pave the way for strategies to control pathogens, enhance biotechnological processes, and deepen the understanding of dormant states across different life forms.
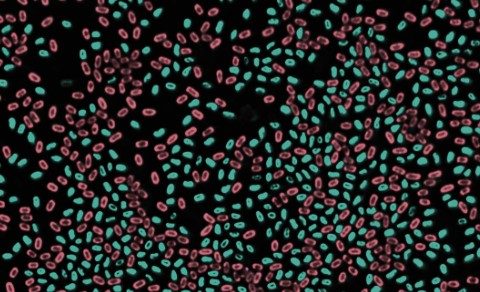
Spores, resilient and protective structures formed by certain microorganisms like bacteria and fungi, serve as a survival mechanism against adverse conditions. Bacterial spores are among the longest-living cellular forms on Earth, with reports evoking their revival following millions of years of dormancy. They encapsulate the organism’s genetic material and essential components, remaining inactive until conditions become favorable for germination. Comprehending spores is crucial in various fields, offering insights into survival strategies and potential applications in microbiology, agriculture, and biotechnology.
Upon emergence from dormancy, the RNA polymerase inside these spores promptly initiates the copying of vital genetic instructions. It recruits necessary components for transcription, such as sigma factors, swiftly activating essential genes necessary for cellular functions. The study also observed a similar process in disease-causing bacteria that form spores, suggesting a common strategy among various organisms to reinitiate functions after dormancy.
“Our research suggests that the structure of the spore chromosome is designed to uphold a blueprint for gene activity by pausing RNA polymerase, in a standby mode, ready to resume gene expression when conditions favor revival,” Ben-Yehuda says. “This mechanism’s relevance might extend beyond bacteria, offering valuable insights into maintaining enduring gene activity plans across various organisms that undergo dormant life stages.”
Researchers:
Bing Zhou1, Yifei Xiong1, Yuval Nevo2, Tamar Kahan3, Oren Yakovian1, Sima Alon1, Saurabh Bhattacharya1, Ilan Rosenshine1, Lior Sinai1, Sigal Ben-Yehuda1
Institutions:
1) Department of Microbiology and Molecular Genetics, Institute for Medical Research Israel-Canada, The Hebrew University-Hadassah Medical School
2) Info-CORE, Bioinformatics Unit of the I-CORE Computation Center at the Hebrew University of Jerusalem
3) Bioinformatics Unit, Faculty of Medicine, The Hebrew University of Jerusalem
Related articles
How Bacteria Outsmart the Immune System: Two-Pronged Strategy Revealed
April 1, 2026 – Researchers at the Hebrew University of Jerusalem (HU) uncovered how a disease-causing bacterium uses a single protein to interfere with the body’s defenses in more than one way, offering a clearer picture of how infections take hold at the cellular level. The study was
New AI Algorithm Could Speed Rare Disease Diagnoses
April 1, 2026 – A new AI algorithm can dramatically hasten the search for the genetic causes of rare diseases, a process that often takes years and often ends without a diagnosis. According to the study published in Genetics Medicine and led by Dr. Christina Canavati and Prof. Yuval Tabach
New Web-based Platform Turning Indoor Green Walls into Smart, Living Systems
April 1, 2026 – Step into a modern office tower or hospital, and the air you breathe is often carefully engineered, filtered, circulated, and cooled at a high energy cost. Now imagine those same spaces quietly breathing on their own, supported by living walls of plants that not only beautify